
Contact
- (210) 567-0884
- xiangy@uthscsa.edu
Department
Microbiology, Immunology & Molecular GeneticsXiang, Yan, Ph.D.
Professor
Personal Statement:
The main focus of Dr. Xiang’s laboratory is on the mechanisms of antiviral immunity and viral strategies of immune evasion. Dr. Xiang is an experienced investigator with over 25 years of experience in various aspects of virology including viral replication, immune evasion, and pathogenesis. The research approaches are distinctly multi-disciplinary, engaging the collaborations with cancer biologists, immunologists and structural biologists.
Education
Ph.D., Virology/Biochemistry at Case Western Reserve University, Cleveland, Ohio
Postdoctoral Training: Laboratory of Viral Diseases, National Institute of Allergy and Infectious diseases, National Institutes of Health
Research
Research interests: The primary interest of Dr. Xiang’s laboratory is host-pathogen interactions, with poxviruses as the model systems. Poxviruses include some dangerous emerging or re-emerging pathogens as well as some promising vaccine vectors for infectious diseases and cancers. The main research projects are: 1. Mechanisms of antiviral immunity and viral strategies of immune evasion. Viruses have evolved ingenious strategies to evade host antiviral immunity. Uncovering these strategies will not only provide insights on viral pathogenesis but also could reveal important aspects of host immunity. Poxviruses are unique among viruses in that they encode a large number of proteins that are dedicated to evading host immune responses. These proteins include secreted inhibitors of cytokines as well as intracellular inhibitors of immune signaling or antiviral factors. Xiang lab has studied extensively a secreted interleukin-18 binding protein (IL-18BP) from poxviruses and their mammalian hosts. Interleukin-18 (IL-18) is a proinflammatory cytokine that can enhance both the innate and acquired immunity. IL-18 protects against microbial infection and tumors. Excessive IL-18 activities, however, are associated with autoimmune and inflammatory diseases. The collaborative work from Xiang lab and Deng lab at Oklahoma State University has elucidated the mechanism by which IL-18BPs bind and inhibit IL-18. The detailed structural knowledge has also been used in developing small molecule inhibitors of IL-18, which could potentially be useful for treating inflammatory diseases. 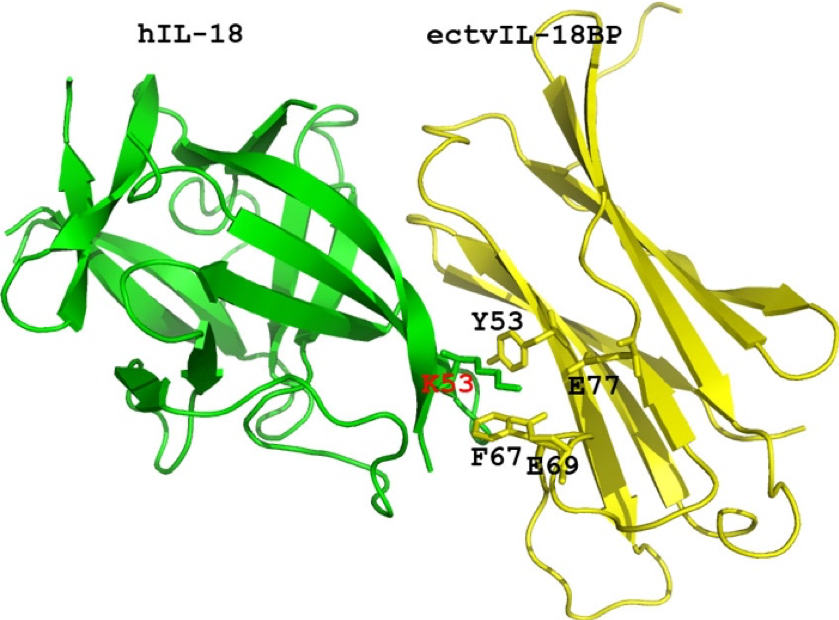 Xiang lab has also studied extensively viral host-range determinants and their specific antagonism of host restriction factors. The interferon (IFN) system is the first line of host defense against intracellular pathogens, particularly viruses. IFNs induce the expression of hundreds of interferon-stimulated genes (ISGs), some of which are antiviral factors that restrict viral replication. To successfully replicate in the cells, viruses have to overcome the formidable barriers posed by the IFN system, particularly the restriction factors. Research from Xiang lab recently showed that a paralogous pair of ISGs, SAMD9 and SAMD9L, form a critical host barrier against poxvirus Infection. Poxviruses, in turn, employ two structurally distinct classes of inhibitors to antagonize SAMD9 and SAMD9L. The outcome of this genetic conflict between poxviruses and their hosts is a major determinant for poxvirus host range. SAMD9 and SAMD9L are tumor suppressors, and their mutations are associated with cancers or developmental disorders.
Xiang lab has also studied extensively viral host-range determinants and their specific antagonism of host restriction factors. The interferon (IFN) system is the first line of host defense against intracellular pathogens, particularly viruses. IFNs induce the expression of hundreds of interferon-stimulated genes (ISGs), some of which are antiviral factors that restrict viral replication. To successfully replicate in the cells, viruses have to overcome the formidable barriers posed by the IFN system, particularly the restriction factors. Research from Xiang lab recently showed that a paralogous pair of ISGs, SAMD9 and SAMD9L, form a critical host barrier against poxvirus Infection. Poxviruses, in turn, employ two structurally distinct classes of inhibitors to antagonize SAMD9 and SAMD9L. The outcome of this genetic conflict between poxviruses and their hosts is a major determinant for poxvirus host range. SAMD9 and SAMD9L are tumor suppressors, and their mutations are associated with cancers or developmental disorders. 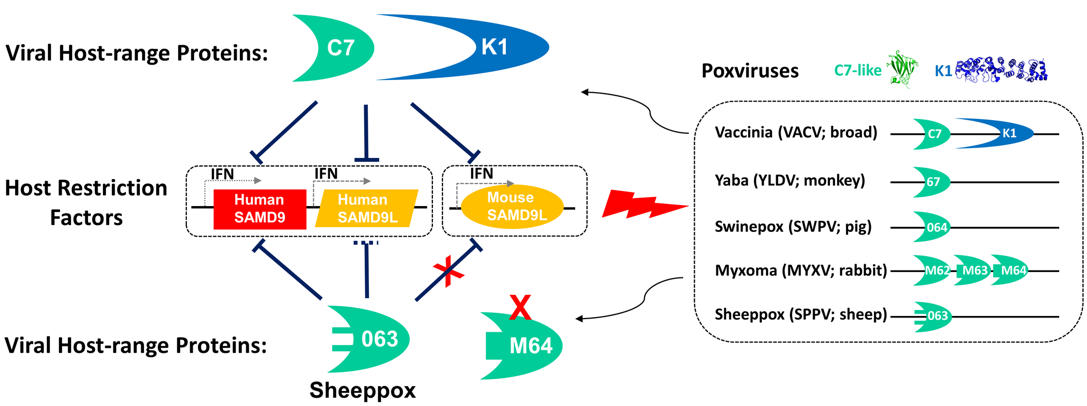 2. Viral assembly mechanism. Viruses, as obligate intracellular parasites, have evolved strategies to manipulate the cellular membranes for entry, genome replication, virion production, and exit. Enveloped viruses typically acquire their outer lipid bilayer by budding from cellular membranes. Poxviruses, however, are unusual in that their primary envelope is not acquired by budding but through extending of open-ended crescent membranes. The origin and biogenesis of the crescent membranes have puzzled virologists for over half a century, albeit recent studies suggest that the crescents may derive from the endoplasmic reticulum (ER). Five viral proteins, conserved in all vertebrate poxviruses and collectively termed viral membrane assembly proteins (VMAPs), have been found to be essential for the biogenesis of crescent membranes. The A6 protein of vaccinia virus (VACV) is the largest VMAP, which Xiang lab discovered and has continued to study its mechanism of action. Uncovering the mechanism by which poxviruses acquire the envelope will not only reveal key viral replication steps for developing antivirals but also provide mechanistic insights on fundamental cellular processes.
2. Viral assembly mechanism. Viruses, as obligate intracellular parasites, have evolved strategies to manipulate the cellular membranes for entry, genome replication, virion production, and exit. Enveloped viruses typically acquire their outer lipid bilayer by budding from cellular membranes. Poxviruses, however, are unusual in that their primary envelope is not acquired by budding but through extending of open-ended crescent membranes. The origin and biogenesis of the crescent membranes have puzzled virologists for over half a century, albeit recent studies suggest that the crescents may derive from the endoplasmic reticulum (ER). Five viral proteins, conserved in all vertebrate poxviruses and collectively termed viral membrane assembly proteins (VMAPs), have been found to be essential for the biogenesis of crescent membranes. The A6 protein of vaccinia virus (VACV) is the largest VMAP, which Xiang lab discovered and has continued to study its mechanism of action. Uncovering the mechanism by which poxviruses acquire the envelope will not only reveal key viral replication steps for developing antivirals but also provide mechanistic insights on fundamental cellular processes. 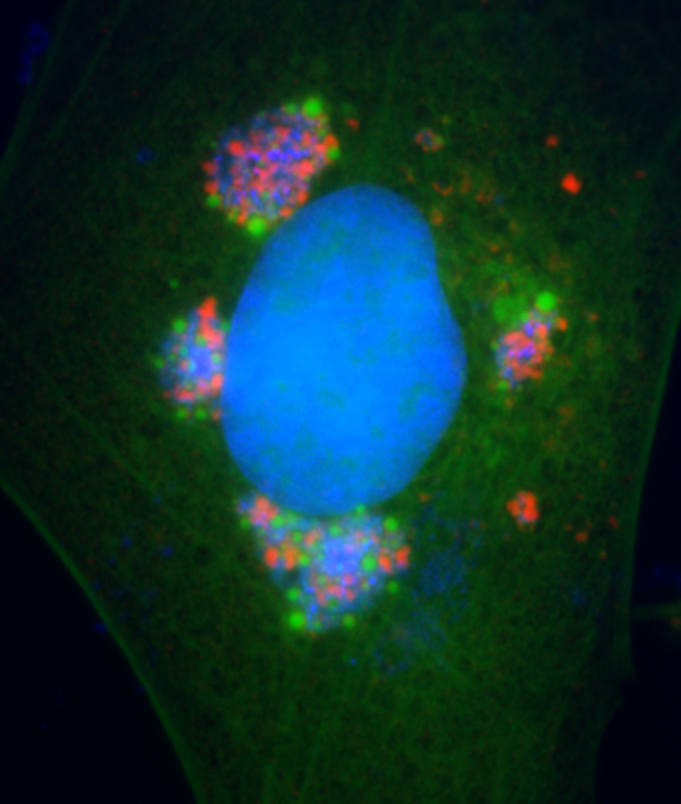 3. Antibody response to vaccination. Vaccinia virus, the live vaccine for smallpox, is one of the most successful vaccines in human history but presents a level of risk that has become unacceptable for the current population. Studying the immune protection mechanism of smallpox vaccine is important for understanding the basic principle of successful vaccines and the development of next generation, safer vaccines. Xiang lab has studied antibody targets in smallpox vaccine by comprehensively characterizing monoclonal antibodies against vaccinia antigens. The knowledge has also been used to optimize vaccinia-based vaccines against other infectious diseases.
3. Antibody response to vaccination. Vaccinia virus, the live vaccine for smallpox, is one of the most successful vaccines in human history but presents a level of risk that has become unacceptable for the current population. Studying the immune protection mechanism of smallpox vaccine is important for understanding the basic principle of successful vaccines and the development of next generation, safer vaccines. Xiang lab has studied antibody targets in smallpox vaccine by comprehensively characterizing monoclonal antibodies against vaccinia antigens. The knowledge has also been used to optimize vaccinia-based vaccines against other infectious diseases. 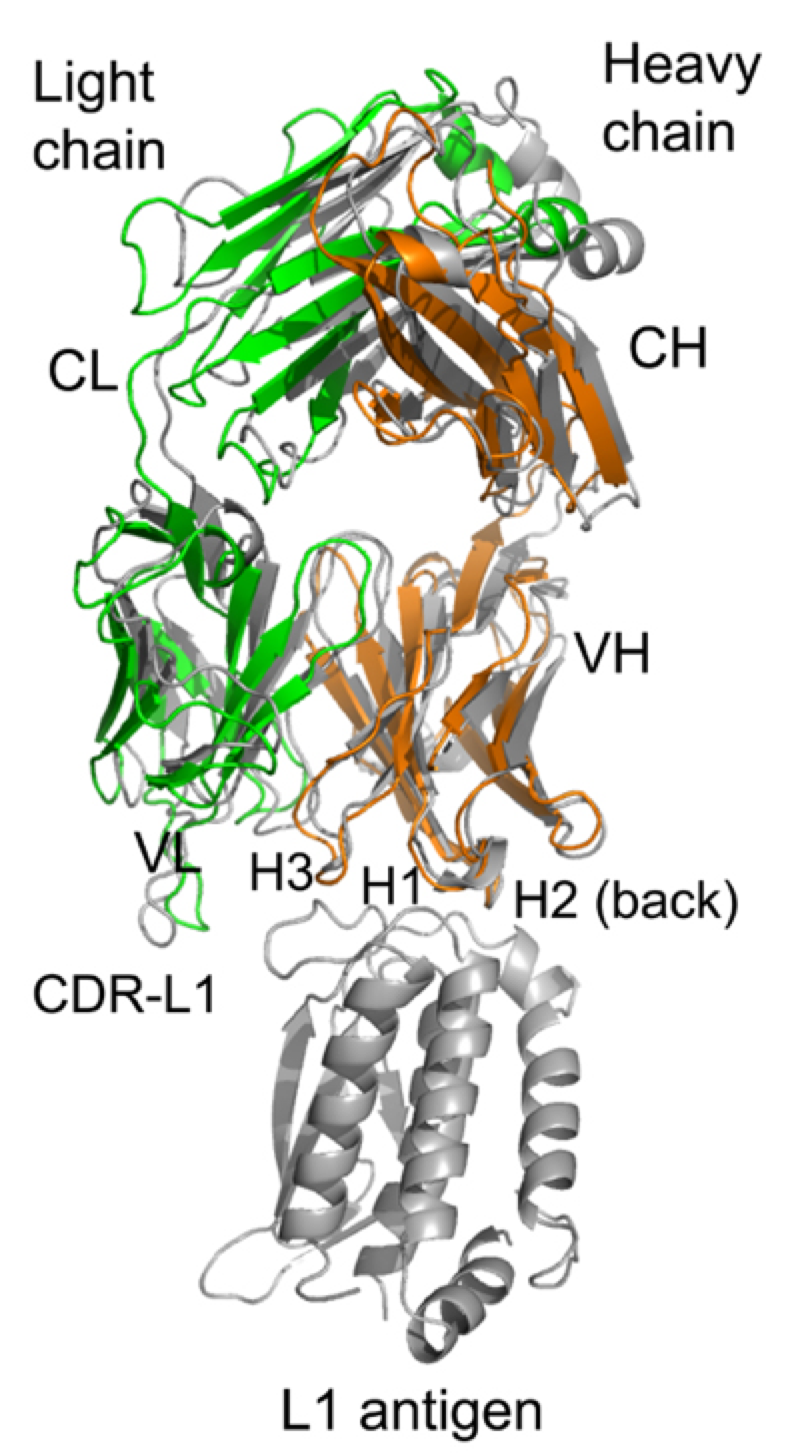
Awards & Accomplishments
Lab Members

| Xiang Lab Members | Name | Position |
|---|---|---|
 | Fushun Zhang, PhD | Research Scientist |
 | Li He, PhD | Postdoctoral Fellow |
 | Marisol Morales | PhD Student |
 | Daniel Sauls | PhD Student |
 | Juan Garcia, Jr. | PhD Student-Co-mentored by Na Xiong, PhD |
Publications
Complete listing of publications: https://www.ncbi.nlm.nih.gov/myncbi/yan.xiang.1/bibliography/public/
- Xiang Y, White A. Monkeypox Virus Emerges from The Shadow of Its More Infamous Cousin: Family Biology Matters. Emerg Microbes Infect. 2022 Jun 24;:1-14. doi: 10.1080/22221751.2022.2095309. [Epub ahead of print] PubMed PMID: 35751396.
- Kitamura N, Sacco MD, Ma C, Hu Y, Townsend JA, Meng X, Zhang F, Zhang X, Ba M, Szeto T, Kukuljac A, Marty MT, Schultz D, Cherry S, Xiang Y, Chen Y, Wang J. Expedited Approach toward the Rational Design of Noncovalent SARS-CoV-2 Main Protease Inhibitors. J Med Chem. 2022 Feb 24;65(4):2848-2865. doi: 10.1021/acs.jmedchem.1c00509. Epub 2021 Apr 23. PubMed PMID: 33891389; PubMed Central PMCID: PMC8536799.
- Peng S, Meng X, Zhang F, Pathak PK, Chaturvedi J, Coronado J, Morales M, Mao Y, Qian SB, Deng J, Xiang Y. Structure and function of an effector domain in antiviral factors and tumor suppressors SAMD9 and SAMD9L. Proc Natl Acad Sci U S A. 2022 Jan 25;119(4). doi: 10.1073/pnas.2116550119. PubMed PMID: 35046037; PubMed Central PMCID: PMC8795524.
- Ma C, Xia Z, Sacco MD, Hu Y, Townsend JA, Meng X, Choza J, Tan H, Jang J, Gongora MV, Zhang X, Zhang F, Xiang Y, Marty MT, Chen Y, Wang J. Discovery of Di- and Trihaloacetamides as Covalent SARS-CoV-2 Main Protease Inhibitors with High Target Specificity. J Am Chem Soc. 2021 Dec 15;143(49):20697-20709. doi: 10.1021/jacs.1c08060. Epub 2021 Dec 3. PubMed PMID: 34860011; PubMed Central PMCID: PMC8672434.
- Hu Y, Meng X, Zhang F, Xiang Y, Wang J. The in vitroantiviral activity of lactoferrin against common human coronaviruses and SARS-CoV-2 is mediated by targeting the heparan sulfate co-receptor. Emerg Microbes Infect. 2021 Dec;10(1):317-330. doi: 10.1080/22221751.2021.1888660. PubMed PMID: 33560940; PubMed Central PMCID: PMC7919907.
- Seifert M, Bera SC, van Nies P, Kirchdoerfer RN, Shannon A, Le TT, Meng X, Xia H, Wood JM, Harris LD, Papini FS, Arnold JJ, Almo S, Grove TL, Shi PY, Xiang Y, Canard B, Depken M, Cameron CE, Dulin D. Inhibition of SARS-CoV-2 polymerase by nucleotide analogs from a single-molecule perspective. 2021 Oct 7;10. doi: 10.7554/eLife.70968. PubMed PMID: 34617885; PubMed Central PMCID: PMC8497053.
- Cantwell AM, Singh H, Platt M, Yu Y, Lin YH, Ikeno Y, Hubbard G, Xiang Y, Gonzalez-Juarbe N, Dube PH. Kinetic Multi-omic Analysis of Responses to SARS-CoV-2 Infection in a Model of Severe COVID-19. J Virol. 2021 Sep 27;95(20):e0101021. doi: 10.1128/JVI.01010-21. Epub 2021 Jul 28. PubMed PMID: 34319784; PubMed Central PMCID: PMC8475517.
- Xia Z, Sacco M, Hu Y, Ma C, Meng X, Zhang F, Szeto T, Xiang Y, Chen Y, Wang J. Rational Design of Hybrid SARS-CoV-2 Main Protease Inhibitors Guided by the Superimposed Cocrystal Structures with the Peptidomimetic Inhibitors GC-376, Telaprevir, and Boceprevir. ACS Pharmacol Transl Sci. 2021 Aug 13;4(4):1408-1421. doi: 10.1021/acsptsci.1c00099. eCollection 2021 Aug 13. PubMed PMID: 34414360; PubMed Central PMCID: PMC8204911.
- Ma C, Sacco MD, Xia Z, Lambrinidis G, Townsend JA, Hu Y, Meng X, Szeto T, Ba M, Zhang X, Gongora M, Zhang F, Marty MT, Xiang Y, Kolocouris A, Chen Y, Wang J. Discovery of SARS-CoV-2 Papain-like Protease Inhibitors through a Combination of High-Throughput Screening and a FlipGFP-Based Reporter Assay. ACS Cent Sci. 2021 Jul 28;7(7):1245-1260. doi: 10.1021/acscentsci.1c00519. Epub 2021 Jun 18. PubMed PMID: 34341772; PubMed Central PMCID: PMC8265724.
- Ma H, Zeng W, Meng X, Huang X, Yang Y, Zhao D, Zhou P, Wang X, Zhao C, Sun Y, Wang P, Ou H, Hu X, Xiang Y, Jin T. Potent Neutralization of SARS-CoV-2 by Hetero-bivalent Alpaca Nanobodies Targeting the Spike Receptor-Binding Domain. J Virol. 2021 Mar 3;. doi: 10.1128/JVI.02438-20. [Epub ahead of print] PubMed PMID: 33658349; PubMed Central PMCID: PMC8139655.
- Zeng W, Ma H, Ding C, Yang Y, Sun Y, Huang X, He W, Xiang Y, Gao Y, Jin T. Characterization of SARS-CoV-2-specific antibodies in COVID-19 patients reveals highly potent neutralizing IgA. Signal Transduct Target Ther. 2021 Jan 29;6(1):35. doi: 10.1038/s41392-021-00478-7. PubMed PMID: 33514692; PubMed Central PMCID: PMC7844101.
- Sacco MD, Ma C, Lagarias P, Gao A, Townsend JA, Meng X, Dube P, Zhang X, Hu Y, Kitamura N, Hurst B, Tarbet B, Marty MT, Kolocouris A, Xiang Y, Chen Y, Wang J. Structure and inhibition of the SARS-CoV-2 main protease reveal strategy for developing dual inhibitors against Mproand cathepsin L. Sci Adv. 2020 Dec;6(50). doi: 10.1126/sciadv.abe0751. Print 2020 Dec. PubMed PMID: 33158912; PubMed Central PMCID: PMC7725459.
- She L, Alanazi HH, Yan L, Brooks EG, Dube PH, Xiang Y, Zhang F, Sun Y, Liu Y, Zhang X, Li XD. Sensing and signaling of immunogenic extracellular RNAs restrain group 2 innate lymphoid cell-driven acute lung inflammation and airway hyperresponsiveness. PLoS One. 2020;15(7):e0236744. doi: 10.1371/journal.pone.0236744. eCollection 2020. PubMed PMID: 32730309; PubMed Central PMCID: PMC7392318.
- Riad S, Xiang Y, AlDaif B, Mercer AA, Fleming SB. Rescue of a Vaccinia Virus Mutant Lacking IFN Resistance Genes K1L and C7L by the ParapoxvirusOrf Virus. Front Microbiol. 2020;11:1797. doi: 10.3389/fmicb.2020.01797. eCollection 2020. PubMed PMID: 32903701; PubMed Central PMCID: PMC7438785.
- Zhang F, Meng X, Townsend MB, Satheshkumar PS, Xiang Y. Identification of CP77 as the Third Orthopoxvirus SAMD9 and SAMD9L Inhibitor with Unique Specificity for a Rodent SAMD9L. J Virol. 2019 Jun 15;93(12). doi: 10.1128/JVI.00225-19. Print 2019 Jun 15. PubMed PMID: 30918078; PubMed Central PMCID: PMC6613757.
- Meng X, Xiang Y. RNA granules associated with SAMD9-mediated poxvirus restriction are similar to antiviral granules in composition but do not require TIA1 for poxvirus restriction. 2019 Mar;529:16-22. doi: 10.1016/j.virol.2019.01.007. Epub 2019 Jan 8. PubMed PMID: 30641480; PubMed Central PMCID: PMC6382536.
- Pathak PK, Peng S, Meng X, Han Y, Zhang B, Zhang F, Xiang Y, Deng J. Structure of a lipid-bound viral membrane assembly protein reveals a modality for enclosing the lipid bilayer. Proc Natl Acad Sci U S A. 2018 Jul 3;115(27):7028-7032. doi: 10.1073/pnas.1805855115. Epub 2018 Jun 18. PubMed PMID: 29915071; PubMed Central PMCID: PMC6142198.
- Meng X, Kaever T, Yan B, Traktman P, Zajonc DM, Peters B, Crotty S, Xiang Y. Characterization of murine antibody responses to vaccinia virus envelope protein A14 reveals an immunodominant antigen lacking of effective neutralization targets. 2018 May;518:284-292. doi: 10.1016/j.virol.2018.03.005. Epub 2018 Mar 17. PubMed PMID: 29558682; PubMed Central PMCID: PMC5911218.
- Meng X, Zhang F, Yan B, Si C, Honda H, Nagamachi A, Sun LZ, Xiang Y. A paralogous pair of mammalian host restriction factors form a critical host barrier against poxvirus infection. PLoS Pathog. 2018 Feb;14(2):e1006884. doi: 10.1371/journal.ppat.1006884. eCollection 2018 Feb. PubMed PMID: 29447249; PubMed Central PMCID: PMC5831749.
- Matho MH, Schlossman A, Gilchuk IM, Miller G, Mikulski Z, Hupfer M, Wang J, Bitra A, Meng X, Xiang Y, Kaever T, Doukov T, Ley K, Crotty S, Peters B, Hsieh-Wilson LC, Crowe JE Jr, Zajonc DM. Structure-function characterization of three human antibodies targeting the vaccinia virus adhesion molecule D8. J Biol Chem. 2018 Jan 5;293(1):390-401. doi: 10.1074/jbc.M117.814541. Epub 2017 Nov 9. PubMed PMID: 29123031; PubMed Central PMCID: PMC5766908.
- Weisberg AS, Maruri-Avidal L, Bisht H, Hansen BT, Schwartz CL, Fischer ER, Meng X, Xiang Y, Moss B. Enigmatic origin of the poxvirus membrane from the endoplasmic reticulum shown by 3D imaging of vaccinia virus assembly mutants. Proc Natl Acad Sci U S A. 2017 Dec 19;114(51):E11001-E11009. doi: 10.1073/pnas.1716255114. Epub 2017 Dec 4. PubMed PMID: 29203656; PubMed Central PMCID: PMC5754806.
- Zaitseva M, Thomas A, Meseda CA, Cheung CYK, Diaz CG, Xiang Y, Crotty S, Golding H. Development of an animal model of progressive vaccinia in nu/nu mice and the use of bioluminescence imaging for assessment of the efficacy of monoclonal antibodies against vaccinial B5 and L1 proteins. Antiviral Res. 2017 Aug;144:8-20. doi: 10.1016/j.antiviral.2017.05.002. Epub 2017 May 8. PubMed PMID: 28495463.
- Meng X, Rose L, Han Y, Deng J, Xiang Y. Vaccinia Virus A6 Is a Two-Domain Protein Requiring a Cognate N-Terminal Domain for Full Viral Membrane Assembly Activity. J Virol. 2017 May 15;91(10). doi: 10.1128/JVI.02405-16. Print 2017 May 15. PubMed PMID: 28275183; PubMed Central PMCID: PMC5411583.
- Krumm B, Meng X, Xiang Y, Deng J. Identification of small molecule inhibitors of Interleukin-18. Sci Rep. 2017 Mar 28;7(1):483. doi: 10.1038/s41598-017-00532-x. PubMed PMID: 28352119; PubMed Central PMCID: PMC5428663.

